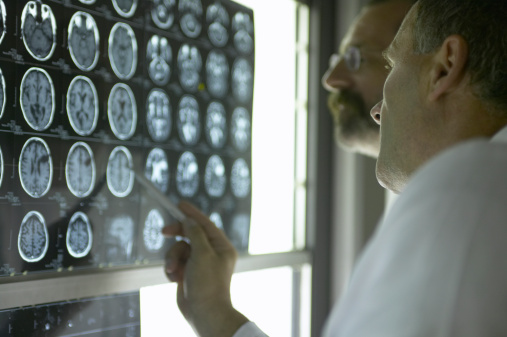
Discovery of new repeat expansion disorders in ataxia and beyond
Date: November 2023
Prepared by SIC Member: Katja Lohmann, PhD
Authors: Melanie Bahlo, PhD, AM, FAHMS; Mathilde Renaud, MD, PhD; Stephan Züchner, MD, PhD, FAAN
Editor: Lorraine Kalia, MD, PhD
Introduction
Late-onset cerebellar ataxia (LOCA) is a group of neurodegenerative disorders that manifest with a progressive cerebellar syndrome after the age of 30 years and are often sporadic (i.e., negative family history). Until recently, there has been an unsatisfactory gap in the genetic diagnosis of patients with LOCA.1 However, in 2019, a new milestone in the field was reached when biallelic repeat expansions (mainly involving AAGGG repeats) were found in an intron of the RFC1 gene, encoding Replication Factor C subunit. Such expansions undoubtedly cause cerebellar ataxia-neuropathy-vestibular areflexia syndrome (CANVAS) and other types of LOCA, often accompanied by neuropathy and/or bilateral vestibulopathy.2,3 An additional exciting milestone was even more recently achieved when two independent research teams discovered GAA repeat expansions in an intron of the FGF14 gene, encoding Fibroblast Growth Factor 14, as a cause of another LOCA subtype (SCA27B) with a seemingly broader phenotypic spectrum.4,5 These discoveries related to LOCA shed new light on ataxia research. Currently, more than 20% of patients previously diagnosed with idiopathic LOCA receive a genetic diagnosis in some studies,4,6 allowing genetic counseling in family members. Prior to the identification of these two genes with previously unrecognized intronic repeat expansions, many were skeptical that there remained prevalent genetic causes of LOCA to be discovered. While it remains to be seen how these new insights can be translated into better treatments for patients, it is exciting to see the new developments in the genetics of ataxia.
What were the clues that led to the identification of the intronic repeat expansions in RFC1 and FGF14?
Dr. Bahlo:
A constellation of advancements occurred over a few years to make these discoveries possible. These included: 1) cheaper whole genome sequencing, allowing sequencing of multiple individuals from families, and now increasingly cohorts of hundreds or even thousands of individuals for discovery, and 2) the development of new bioinformatics tools that made the discovery of repeat expansions in whole genome (and exome) sequencing data possible.7 However, for both RFC1 and FGF14, there were additional clues that helped to make these discoveries. For RFC1-linked CANVAS, pedigree studies had previously localized the causal variant to a small region of the genome, while for FGF14 there was strong prior evidence since small sequence variants in FGF14 were already known to cause a form of ataxia.8
Ataxias are a highly heterogeneous group of diseases, both clinically and genetically. Currently, the intronic repeat expansions in RFC1 and FGF14 are not detectable with standard gene panels or exome sequencing, thus requiring specific testing. Are there certain clinical features to flag patients for testing?
Dr. Renaud:
The core phenotype of SCA27B (GAA-FGF14 ataxia) consists of a slowly progressive cerebellar syndrome characterized by gait ataxia and cerebellar oculomotor impairment. While most patients present with gait unsteadiness at disease onset, almost half of patients report episodic symptoms such as vertigo and/or dizziness, visual disturbances (diplopia, oscillopsia, blurring), and dysarthria. Episodes can be induced by alcohol, physical activity, or sometimes caffeine. More than half of patients with SCA27B display sensitivity to alcohol, which may trigger episodes of ataxia or dramatically worsen baseline ataxia. Downbeat nystagmus, cerebellar oculomotor signs, impaired visual fixation suppression of the vestibular-ocular reflex, vertiginous symptoms, and visual disturbances frequently co-occur at disease onset. In addition to cerebellar impairment, vestibular hypofunction and afferent sensory defect can be observed in SCA27B. Bilateral vestibulopathy is possible with SCA27B but remains relatively mild, while it is almost always present in RFC1-related CANVAS and disease spectrum. Polyneuropathy is not a core feature of SCA27B, although some patients may develop mild axonal sensory or sensorimotor polyneuropathy. In comparison, motor neuropathy is typically mild or absent in RFC1-related CANVAS. The diagnosis of RFC1-related CANVAS and disease spectrum is highly unlikely in the presence of isolated cerebellar ataxia without sensory neuropathy. Disease progression in SCA27B, averaging 0.23-0.40 SARA points per year, is considerably slower than in other common late-onset genetic ataxias, such as SCA6 (0.80 SARA points per year) and RFC1-related CANVAS (1.30 SARA points per year).
Do you think there are additional repeat expansion disorders to be discovered?
Dr. Bahlo:
Yes, I do think there are more repeat expansions to discover. I believe there are two main areas where there are likely ‘hidden’ repeat expansion disorders. First, repeat expansions are low mutation rate events that are often only found in specific human populations or among people of particular ancestries. Some human populations are highly underrepresented in our genome sequencing efforts and are also underrepresented for repeat expansion compositions. It will be fruitful to look into such populations for novel repeat expansions. The second area is the detection of the more challenging to identify repeat expansions with ‘incomplete penetrance,’ i.e. where there is an increased risk of disease rather than certainty of getting the disease. The discovery of these types of repeat expansions requires larger cohorts and different statistical methods. Repeat expansions in FGF14 are an example of such variants, but the repeat is still highly penetrant.
Could intronic repeat expansions also have a role in other neurodegenerative disorders, such as Parkinson´s disease?
Dr. Renaud:
Yes, it is conceivable that intronic repeat expansions have a role in other neurodegenerative disorders, such as Parkinson´s disease. New pathogenic intronic repeat expansions will likely be discovered in the near future thanks to the widespread implementation of long-read sequencing and advanced bioinformatics tools. Detailed deep phenotyping of patient cohorts is, however, an essential prerequisite to increase the chances of finding new pathogenic repeat expansions in neurodegenerative diseases.
What is known about the functional consequences of the intronic repeat expansions and the role of the specific motif?
Dr. Züchner:
We are still only beginning to understand the molecular mechanisms behind RFC1 and FGF14 repeat expansion disorders. The novel repeat expansions in RFC1 and FGF14 are both located within the non-coding intronic region and do not have an obvious impact on the protein sequence. Further, interruptions of the repeated sequence seem to be protective. There is published evidence of a reduced gene expression that is likely transcript-specific.4 It is also interesting that FGF14 is not widely expressed; thus, its cell-specific roles in certain neuron populations are at the core of the mechanism of action. Beyond these loss-of-function hypotheses, there might be other mechanisms, such as RAN (Repeat Associated Non-AUG) translation, that would lead in fact to a toxic gain-of-function. We can expect many interesting functional studies on these genes and their pathways in the coming years.
The role of the repeat motifs is very much appreciated. The broader adaption of long-read sequencing methods allows for dissecting the composition of very long repeat stretches with excellent fidelity. It turns out that a sizable spectrum of repeat motifs, interruptions, and periodicity can be observed in the general population and in patients. The pathogenicity of each of these motifs is currently intensely studied and will inform diagnostic assays. Finally, exciting new findings published by Pellerin et al.9, indicate that a specific 17bp sequence motif located immediately 5’ of the FGF14 GAA repeat determines the meiotic expansion stability of the locus. The discovery of such a robust predicting motif is a first in the field of repeat expansions.
What are the next steps to further improve genetic testing for patients with ataxia?
Dr. Bahlo:
We need to implement our current set of bioinformatic tools as very potent screening tools in clinical and research cohorts. Precise genotyping is not required for these to have a significant impact in clinical pipelines where they will detect currently missed repeat expansions. This will lead to re-diagnoses for some patients and increased solve rates overall. Several large cohort studies with extensive validation work with traditional methods have shown that the methods are highly sensitive and specific, over 95%, especially for the smaller repeat expansions. Hits should then be validated with the current, repeat expansion specific, gold standard tests, such as repeat-primed PCR and Southern blots. In the future, these assays will be replaced by deep long-read sequencing, which is already proving to be incisive in characterizing the increasingly heterogeneous complex motifs found.
How can these insights be translated into new treatments?
Dr. Züchner:
Gene-specific treatments, such as gene editing or small molecules targeted to specific repeat motifs, will very soon be tested in RFC1- and FGF14-related diseases. To be effective, it will be essential to understand whether loss- and/or gain-of-function mechanisms are at play. This will very likely require well-characterized animal models for these diseases, which are being worked on in our lab and by others. For FGF14, a long-existing treatment for episodic ataxia, 4-amino-pyridine, has recently been explored10, and a larger trial will possibly follow. The fact that a potassium channel blocker can modify the outcome of FGF14-related ataxia points to a mechanistic link that can likely be further exploited in the future. In summary, there is reason for optimism of timely translational studies for these diseases.
Conclusions
The recent breakthrough in the genetics of LOCA is the identification of two novel repeat expansion disorders that account for a considerable number of patients. As pointed out by the interviewed experts, more insights are going to be expected in terms of a better molecular understanding of these two repeat expansion disorders, but likewise for novel ones and certainly also beyond the phenotype of LOCA. Research into how to translate these novel findings into better treatments for affected individuals is already underway. Thus, the next steps in this rapidly moving field include: 1) search for novel repeat expansions by including underrepresented populations and considering reduced penetrance, 2) further elucidating the phenotypic and genotypic spectrum of RFC1- and FGF14-related diseases taking the composition of the repeat motifs and other variants in these and other genes into account, and 3) translation of the findings into improved therapies. Certainly, the human genome with its hundreds of thousands of repeat sequences is very complex, and we are still far away from understanding the complete picture, but the future promises even deeper insights.
References:
1. van Gaalen J, van de Warrenburg BP. A practical approach to late-onset cerebellar ataxia: putting the disorder with lack of order into order. Pract Neurol 2012;12:14-24.
2. Cortese A, Simone R, Sullivan R, et al. Biallelic expansion of an intronic repeat in RFC1 is a common cause of late-onset ataxia. Nat Genet 2019;51:649-658.
3. Rafehi H, Szmulewicz DJ, Bennett MF, et al. Bioinformatics-Based Identification of Expanded Repeats: A Non-reference Intronic Pentamer Expansion in RFC1 Causes CANVAS. Am J Hum Genet 2019;105:151-165.
4. Pellerin D, Danzi MC, Wilke C, et al. Deep Intronic FGF14 GAA Repeat Expansion in Late-Onset Cerebellar Ataxia. N Engl J Med 2023;388:128-141.
5. Rafehi H, Read J, Szmulewicz DJ, et al. An intronic GAA repeat expansion in FGF14 causes the autosomal-dominant adult-onset ataxia SCA50/ATX-FGF14. Am J Hum Genet 2023;110:105-119.
6. Iruzubieta P, Pellerin D, Bergareche A, et al. Frequency and phenotypic spectrum of spinocerebellar ataxia 27B and other genetic ataxias in a Spanish cohort of late-onset cerebellar ataxia. Eur J Neurol 2023.
7. Bahlo M, Bennett MF, Degorski P, Tankard RM, Delatycki MB, Lockhart PJ. Recent advances in the detection of repeat expansions with short-read next-generation sequencing. F1000Res 2018;7.
8. van Swieten JC, Brusse E, de Graaf BM, et al. A mutation in the fibroblast growth factor 14 gene is associated with autosomal dominant cerebellar ataxia [corrected]. Am J Hum Genet 2003;72:191-199.
9. Pellerin D, Gobbo GD, Couse M, et al. A common flanking variant is associated with enhanced meiotic stability of the FGF14 -SCA27B locus. bioRxiv 2023.
10. Wilke C, Pellerin D, Mengel D, et al. GAA-FGF14 ataxia (SCA27B): phenotypic profile, natural history progression and 4-aminopyridine treatment response. Brain 2023;146:4144-4157.