VOLUME 26, ISSUE 3 • SEPTEMBER 2022 Full issue »
VOLUME 26, ISSUE 3 • SEPTEMBER 2022 Full issue »

2022 Updates: Nomenclature of Genetic Movement Disorders
New genes, classification enhancements introduced
Originally, locus symbols (e.g., DYT1) were used to specify chromosomal regions that had been linked to a familial disorder or a specific phenotype with an as yet unknown gene. These symbols were assigned in a chronological, numerical order (e.g., PARK1, PARK2, etc.) and were regularly used in lieu of the name for the condition (e.g., DYT1 dystonia), even when the disease-causing gene (e.g., TOR1A for DYT1) had been identified. This system had a number of weaknesses, e.g., the inability to distinguish disease-causing genes from genetic risk factors, more than one symbol being assigned for the same disorder, unconfirmed associations between a gene or locus and a movement disorder, or designated symbols in the absence of any known locus or gene. 1, 2
Therefore, the International Parkinson and Movement Disorder Society (MDS) Task Force for the Nomenclature of Genetic Movement Disorders was initiated to revise this system. Starting in 2016, the Task Force presented a new system for naming genetically determined movement disorders and provided a criterion-based list of confirmed monogenic movement disorders including parkinsonism, dystonia, cerebellar ataxia, chorea, hereditary spastic paraplegia, paroxysmal movement disorders, myoclonus, primary familial brain calcification, and neurodegeneration with brain iron accumulation. 1, 3, 4 These criteria included a convincing level of evidence for a gene to be disease causing (e.g., reported variants in multiple unrelated affected patients, evidence for segregation or from functional studies and in silico prediction tools), independent confirmation in the literature and a movement disorder as the predominant phenotype linked to the gene.
Since then, next-generation sequencing techniques have led to the identification of a substantial number of new disease-causing genes, which warrant classification using the novel system. The MDS Task Force published an update in 2022 including not only newly identified and confirmed disease-causing genes but also introducing a number of additional changes to the classification system. 5
First, we proposed to dissolve the imaging-based categories of Primary Familial Brain Calcification and Neurodegeneration with Brain Iron Accumulation and to reclassify these genetic conditions according to their predominant phenotype, aligning with the movement phenotypic emphasis in the nomenclature. Still, to highlight the imaging features, the iron accumulation and brain calcification features of these entities are referenced, e.g., PARK-SLC20A2-(PFBC). We further introduced the novel category and prefix of mixed movement disorders (MxMD), which includes conditions linked to more than two prominent movement disorder phenotypes, either co-existing or having been described in an approximately equal number of patients.
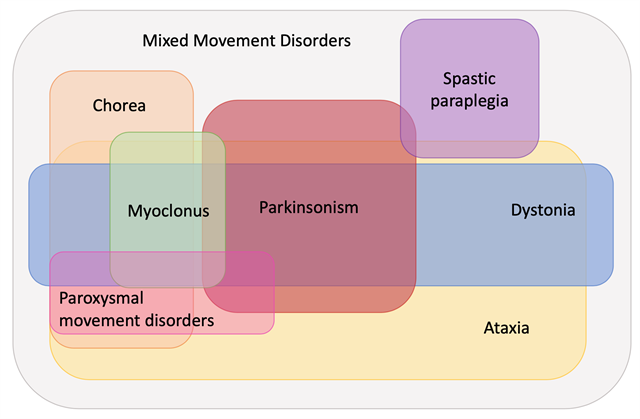
In keeping with the rapid pace of gene discovery, in this update since 2016, we listed a total of 89 newly identified genes that warranted a prefix based on our criteria: 6 genes for parkinsonism, 21 for dystonia, 38 for dominant and recessive ataxia, 5 for chorea, 7 for myoclonus, 13 for spastic paraplegia, 3 for paroxysmal movement disorders, and 6 for mixed movement disorder phenotypes; 10 genes were linked to two phenotypes and have been assigned double prefixes. We also prepared lists of yet unconfirmed gene-disease associations in which the respective movement disorder has only been described in the minority of cases or as a less prominent feature. These lists include another 14 genes for parkinsonism, 12 for dystonia, 96 for dominant and recessive ataxia, 7 for chorea, 13 for myoclonus, and 20 for spastic paraplegia. Each entry is accompanied by a ‘clinical clues’ entry to guide the clinician. It is intended that the Task Force will be engaged with periodic updates at approximately 5-year intervals.
These updated nomenclature lists have also been published on the MDS website:
References
1. Marras C, Lang A, van de Warrenburg BP, et al. Nomenclature of genetic movement disorders: Recommendations of the international Parkinson and movement disorder society task force. Mov Disord 2016;31(4):436-457.
2. Marras C, Lohmann K, Lang A, Klein C. Fixing the broken system of genetic locus symbols: Parkinson disease and dystonia as examples. Neurology 2012;78(13):1016-1024.
3. Rossi M, Anheim M, Durr A, et al. The genetic nomenclature of recessive cerebellar ataxias. Mov Disord 2018;33(7):1056-1076.
4. van der Veen S, Zutt R, Klein C, et al. Nomenclature of Genetically Determined Myoclonus Syndromes: Recommendations of the International Parkinson and Movement Disorder Society Task Force. Mov Disord 2019;34(11):1602-1613.
5. Lange LM, Gonzalez-Latapi P, Rajalingam R, et al. Nomenclature of Genetic Movement Disorders: Recommendations of the International Parkinson and Movement Disorder Society Task Force - An Update. Mov Disord 2022;37(5):905-935.
Read more Moving Along: